Research
Biosensors enable the selective and sensitive detection of biological analytes and biological processes. As such, biosensors have an ever-increasing impact in healthcare, food safety and production, and environmental monitoring. In the biomedical context, quickly identifying antimicrobial resistance during bacterial infections or efficiently detecting proteins and signalling molecules (i.e., cytokines) in human samples is crucial for choosing the most effective treatment for patients. Evolution has formed a shear endless pool of proteins, enzymes, signalling molecules and reaction cascades, many of which are yet unknown or poorly characterised.
The current and planned research in the lab can be loosely grouped into three areas: (1) Live-cell biosensing, (2) DNA- and RNA-based in vitro biosensing, and (3) Functional super-resolution microscopy and microfluidics for lab-on-the-chip systems
1. Live-cell biosensing
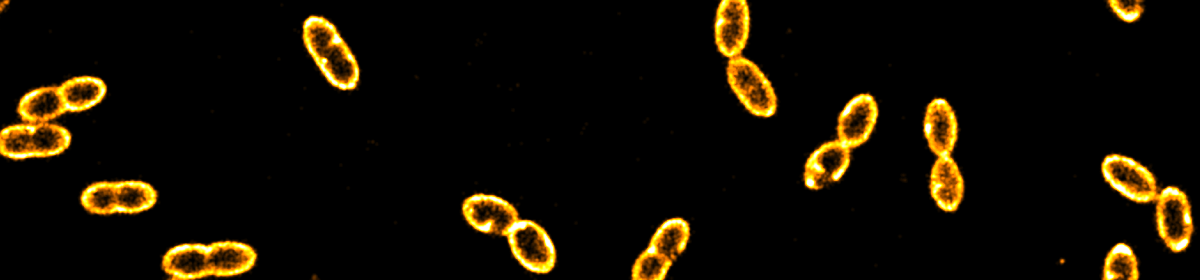
Challenges and research gap. Microbes are indispensable in food production (e.g., flavours, texture, nutrients), biodegradation (e.g., breaking down of organic matter during wastewater treatment), pharmacy (e.g., producing antibiotics), and other industrial applications. In foods, lactic acid bacteria (LAB) starter cultures are essential for food products such as cheese and yoghurt. LAB are also promising candidates for the production of reagents for medical diagnostics and therapeutics (Song et al., 2017). With the current legislation hindering direct genomic engineering, current strategies to improve LAB species are cumbersome and expensive. A promising alternative for the improvement of LAB species is harnessing natural competence, which enables bacteria to gain genetically encoded traits. Natural competence is a result of finely tuned biosensing in which bacteria read out environmental cues to adapt their metabolism and genetic fate. For most LAB species, however, the molecular details and exact pathways of this phenomenon are unknown awaiting exploration. We further work on CRISPR-Cas, a bacterial defence mechanism that protects bacteria against foreign DNA or RNA. CRISPR-Cas utilises short guide RNA as probes to find complementary RNA or DNA sequences and, upon identification of the DNA/RNA as foreign, initiates their cleavage and destruction. As such, CRISPR-Cas has found numerous applications in gene editing and biosensing.
2. DNA- and RNA-based in vitro biosensing
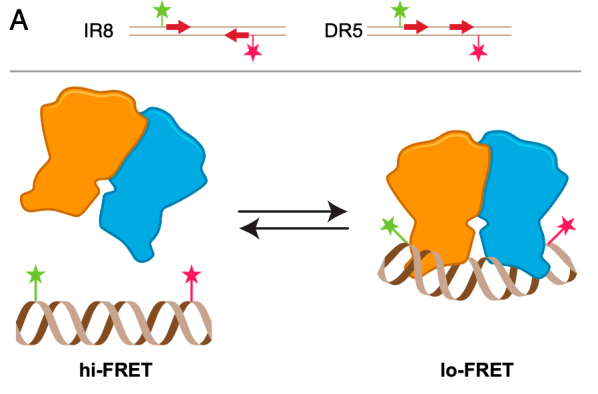
Challenges and research gap. Biosensing relies on the efficient detection of biological processes. One example of a highly customisable scaffold for biosensing is DNA. For instance, DNA can be utilised as a single strand selected to bind target proteins (DNA aptamer) or as a short, double-stranded, one-dimensional DNA oligomer. Additionally, so-called DNA origami, in which a long templating strand is mixed in solution with many short complementary strands, are able to anneal into complex three-dimensional structures. DNA origamis have already been employed as force sensors and even as rotary ratchet motors showcasing their wide versatility. Previously, we used short DNA oligomers to develop assays allowing the detection of specific DNA binding proteins, namely transcription factors and DNA polymerases. We combined the sensors with single-molecule FRET, a technique that allows measuring conformational changes of biomolecules in the nanometre range. One long-standing challenge in the context of biosensors is to develop assays that enable multiplexing and high throughput. In this sense, the use of DNA sequencing microfluidic chips and camera-based real-time sensing will be explored.
3. Super-resolution microscopy and microfluidics
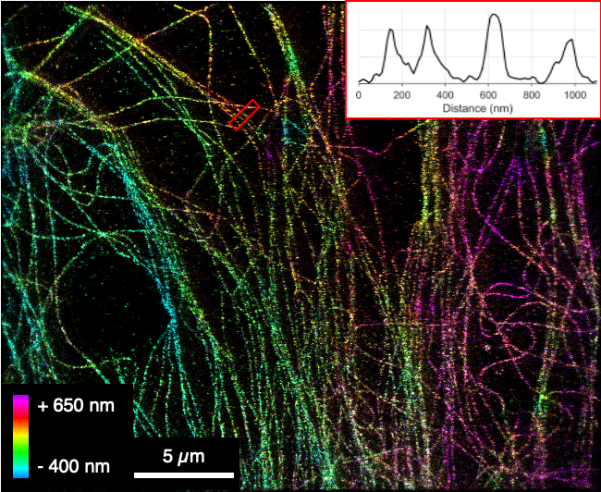
Challenges and research gap. Recent breakthroughs in fluorescence-based microscopy such as MINFLUX, MinSTED, and RESI have pushed the achievable resolution in biomedical imaging to the sub-nanometer regime. Having the technical background and deep understanding of these technologies, our group is renowned for pushing the boundaries of single-molecule detection with a clear focus on simplifying technologies to lower entry barriers via open sourcing of hardware and software. We have been working on specially designed nanofluidic devices that combine the advantages of diffusion-based, confocal microscopy (providing high time resolution for studying fast conformational changes) and camera-based microscopy (with the ability to monitor many individual molecules in parallel). We further develop and apply fluidic devices for the continuous monitoring of bacterial cells. Combined we believe, that these developments are bound to deeply impact the field of biosensors.